Drug Resistant Bacteria — A Research Challenge
There has been much ado about drug resistant bacteria in the news. As a researcher in the field, I thought it would be valuable to outline some of the biochemical background that explains some of the challenges in this field.
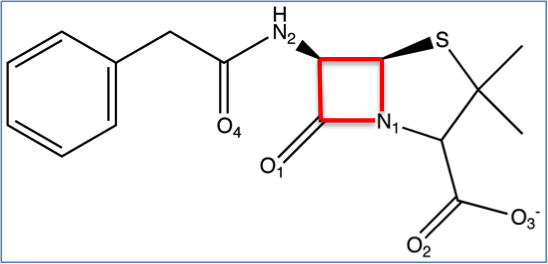
Most bacteria require a cell wall to live. Enzymes named D-alanyl-D-alanine carboxypeptidases (peptidases) are responsible for forming the bacterial cell wall. Beta-lactam antibiotics (i.e. penicllins, cephalosporins, carbapenams, and monobactams) are useful because they inhibit the mechanism peptidases use for bacterial cell wall formation.
However, in response to environmental factors, bacteria have been evolving for millions of years to circumvent these problems. One resistance mechanism is the development of a competing enzyme known as a beta-lactamase. They have a similar binding site therefore compete with peptidases to bind beta-lactam antibiotics. When a beta-lactamase binds a beta-lactam antibiotic it is rendered useless against their intended target, peptidases.

One of the main problems surrounding research today is that beta-lactamases have become better at binding most antibiotics than peptidases. However, this is not the only method of drug resistance, similarly, beta-lactams are not the only class of antibiotics encountering resistance. For example, bacterial mutations can occur making drug resistance a multifaceted problem. This has resulted in a world wide health crisis. Researchers are scrambling to design new antibiotics to overcome the propagating issue.
The term ‘superbug’ is being used to denote multi-drug resistant (MDR) bacteria. MRSA (methicillin-resistant Staphylococcus aureus), a well known super bug, is common in densely populated areas particularly hospitals, prisons, and athletic locker rooms. A great deal of research is underway to develop effective antibiotics for MRSA, a gram-positive bacteria. In the last two decades, only two new classes of antibiotics (daptomycin and linezolid) have been introduced and are gram-positive specific. Despite this development, MRSA is still a prevalent problem.
This leads to another issue, not all bacteria are gram-positive, additionally, some are gram-negative. In general, there has been less research focused on gram-negative bacterial infections. In 2008, a new beta-lactamase was identified in Klebsiella pneumoniae in India now named the New Delhi metallo-beta-lactamase (NDM-1). This enzyme is able to inhibit all beta-lactams except aztreonam, however its mechanisms make it resistant to nearly all antibiotics. NDM-1 is becoming prevalent in a variety of bacterial strains creating significant complications and a new target for researchers.
We are basically in a ‘keeping up with the Joneses’ relationship with bacteria. We develop new drugs and they evolve.
Some hope…
Research is consistently being done to help solve these problems. A common strategy is to combine a beta-lactamase inhibitor with a beta-lactam antibiotic. Therefore the inhibitor prevents the beta-lactamase from interfering with the antibiotic activity of the beta-lactam in peptidases.
Just this week, I have read a couple promising papers about beta-lactamase inhibitors. The group of Christian Melander at NC State University published a study in Medicinal Chemistry Letters that resulted in a promising drug that acts as a partner to an antibiotic. Their drug binds to NDM-1 and allows the antibiotic to inhibit K. pneumoniae growth. At University of South Florida, Yu Chen’s group published in the Journal of Medicinal Chemistry another beta-lactamase inhibitor for the CTX-M class.
Multi-drug resistant bacteria is a world wide problem and nothing short of consistent, collaborative research will allow us to ‘keep up’.





Fascinating.
To illustrate the problem we face in slowing the overuse of antibiotics, which is a little off-topic but not really.
My daughter was pulled from school on Tuesday because they suspected she had pinkeye. She has also has had a URI and a slight cough for the last week and a half. While at the doctor I was informed that she indeed had conjunctivitis (a mild case) and was given drops, so sorted. The doctor, when asked, told me that the cough was allergic rhinitis which made sense because we are all hacking around here.
Anyway, when I told my wife that there was little that could be done about her cold she said that I should have insisted on antibiotics to clear it up. When I explained that it was most likely a virus and that antibiotics could not help she said, and I am not kidding, ”whatever, that’s your opinion.” While my wife is not entirely scientifically literate she is not stupid but she simply could not grok that viruses and bacteria were completely separate things.
I venture to say that, had my wife taken our daughter to the doctor, she would have been taking unneeded antibiotics. I fear that more people are like my wife than like me. While that might make that would less full of assholes it is making staying alive much more challenging.
I don’t get why people ven *want* to take antibiotics unless the absolutely need them. Antibiotics suck. The side-effects suck. I hate them. I had to take a lot last year due to an infection that would not go away, and I do not miss the weird taste in my mouth, the way it made my food taste funny, the way it killed my appetite, the way it made me queasy, the way it gave me a yeast infection…
Ugh.
Nice editing,
She has also has had a URI
She has also had a URI
While that might make that would less full of assholes
While that might make the world less full of assholes
I’m not drunk, I swear.
I don’t get why people ven *want* <— Well, at least you're not typing made-up words for no reason. "ven"? WTF? lol
You address two important points: viruses vs. bacteria and antibiotic overuse.
Hopefully, from the article it is clear that antibiotics are only used to treat bacteria. Bacteria are living organisms therefore something can be done to kill them. Viruses don’t necessarily fall into the classification of living. They have both living and non-living characteristics. It is true that many people ‘want’ them prescribed for symptoms resulting from a virus. However, the antibiotic mechanisms simply will not work on them. Maybe this can help your wife…
I almost included a bit about the overuse of antibiotics, but I am not a MD so I excluded it. In general, doctors I have spoken with are becoming more and more careful when prescribing antibiotics due to the emerging issues discussed here. One would hope that doctors are always using their education vs a parent’s misconception when treating patients.
If I can get her to sit down and listen for a few minutes I believe I can get her to grasp the difference, or at least as well as I understand it, like I said she’s not stupid. Just for fun, after I get her caught up on that I may introduce her to prions; not literally of course. :)
Good coverup. Always got to remember that plausible denial. :)
Although the most shittingly terrifying thing I read recently was about the totally drug resistant TB.
Factory farming uses antibiotics to stop animals getting sick in crowded pens, it would be hard to design a better way to breed drug resistant bacteria. Any new drugs need to be released along with strict guidlines that limit their use, and please, everyone, avoid factory farm products. Thanks for an interesting article.
In our clinical microbiology lab we isolate MRSA from patients’ cultures almost every day. Contrast this with just 10 or 15 years ago when it happened perhaps once a month.
My wife is a physician and she says many patients simply demand antibiotics regardless of whether they are clinically indicated or not. They aren’t satisfied unless they walk out of the office with a prescription no matter how much you try to reason with them. Doctors try to be responsible with prescibing antibiotics but there is a lot of pressure to give in to their patient’s demands.
So if a patient demands OxyContin, do physicians just give in and prescribe it? Seems that would be less dangerous, since painkiller abuse doesn’t have a murderous effect on the entire population like drug-resistant bacteria does.
Well, I didn’t say doctors just give in and prescribe antibiotics to everyone that demands it…just that there is a lot of pressure to do so.
BTW, there are patients (generally refered to as “drug seekers”) who do whatever they can to get prescription narcs like oxycontin. They are very skilled in the arts of deceit and manipulation and are incredibly dilligent. They’ll go from practice to practice, doctor to doctor until they find one naive or inexperienced enough to give them what they want. It’s often hard for doctors to know whether a patient is really in pain or is lying just to get painkillers. It takes a lot of skill and clinical experience to be able to tease apart the half-truths and lies. And I disagree that this would be less dangerous than antibiotics. Much of the narcs obtained in this manner are sold on the street and contribute to the human misery and violence associated with the illegal drug trade.
The heck with TB. How about totally antibiotic resistant gonorrhea? That one is here.